Set up
library (akldogs)library (tidyverse)
── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
✔ dplyr 1.2.0 ✔ readr 2.1.6
✔ forcats 1.0.1 ✔ stringr 1.6.0
✔ ggplot2 4.0.2 ✔ tibble 3.3.1
✔ lubridate 1.9.5 ✔ tidyr 1.3.2
✔ purrr 1.2.1
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errors
library (tidytext)library (fs)library (knitr)library (kableExtra)
Attaching package: 'kableExtra'
The following object is masked from 'package:dplyr':
group_rows
library (patchwork)library (scales)
Attaching package: 'scales'
The following object is masked from 'package:purrr':
discard
The following object is masked from 'package:readr':
col_factor
Attaching package: 'Hmisc'
The following objects are masked from 'package:dplyr':
src, summarize
The following objects are masked from 'package:base':
format.pval, units
Attaching package: 'plotly'
The following object is masked from 'package:Hmisc':
subplot
The following object is masked from 'package:ggplot2':
last_plot
The following object is masked from 'package:stats':
filter
The following object is masked from 'package:graphics':
layout
source ("../R/functions.R" )source ("../R/theme.R" )
<- list.files ("../data" , pattern = "am-reports" , full.names = TRUE )<- report_files |> path_file () |> path_ext_remove () |> str_extract ("[^-]+$" )<- report_files |> set_names (report_names) |> map (read_csv)
Rows: 96 Columns: 3
── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
chr (1): category
dbl (2): year, count
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.
Rows: 48 Columns: 3
── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
chr (1): category
dbl (2): year, count
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.
Rows: 108 Columns: 3
── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
chr (1): category
dbl (2): year, count
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.
Rows: 224 Columns: 3
── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
chr (1): category
dbl (2): year, count
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.
Simplify some column names
<- registration |> rename (suburb = suburb_name)<- rename (rfs, suburb = location_suburb_name)
Auckland Dog Population Overview
As the dog population has increased, rates of registration and desexing have fallen, while rates of menacing dog ownership have increased.
<- $ ownership |> pivot_wider (names_from = category,values_from = count|> arrange (year) |> mutate (prev_value = lag (dogs),change = dogs - prev_value,pc_change = change / dogs,reg_pc = registered / dogs,desex_pc = desexed / dogs,men_pc = menacing / dogs|> select (year, dogs, pc_change: men_pc) |> arrange (desc (year)) |> rename ("Year" = year,"Known Dogs" = dogs,"Pop. Change" = pc_change,"Registered" = reg_pc,"Desexed" = desex_pc,"Menacing" = men_pc|> mutate (across ("Pop. Change" : "Menacing" ,~ scales:: percent (., accuracy = 0.1 )),"Known Dogs" = format (` Known Dogs ` , big.mark = "," )|> kable () |> kable_styling (full_width = FALSE ) Standarise the spelling of primary breeds
<- akldogs:: breeds<- registration |> left_join (breed_lookup, by = c ("animal_breed_description" = "raw_name" )) |> rename (breed_standardised = standardised_name) |> relocate (breed_standardised, .after = animal_breed_description)<- impounds |> left_join (breed_lookup, by = c ("primary_breed" = "raw_name" )) |> rename (breed_standardised = standardised_name) |> relocate (breed_standardised, .after = primary_breed) Investigate breed mismatches between registration and impound data.
The breed recorded in impound data does not match the breed for the dog with the same animal_id in the registration data, in over 6,000 cases. The rest of the analyses in this notebook use the data as provided by Animal Management, but noting this for follow up.
# Unique animals in impound data <- impounds |> select (dog_id) |> unique ()nrow (impounds_unique)
<- inner_join (|> select (animal_id, breed_reg = clean_name),|> select (dog_id, breed_imp = clean_name),by = c ("animal_id" = "dog_id" )|> unique () |> mutate (is_match = (breed_reg == breed_imp))
Warning in inner_join(select(registration, animal_id, breed_reg = clean_name), : Detected an unexpected many-to-many relationship between `x` and `y`.
ℹ Row 13 of `x` matches multiple rows in `y`.
ℹ Row 198 of `y` matches multiple rows in `x`.
ℹ If a many-to-many relationship is expected, set `relationship =
"many-to-many"` to silence this warning.
<- comparison_by_id |> summarise (total_dogs_checked = n (),perfect_matches = sum (is_match),mismatches = sum (! is_match),accuracy_rate = mean (is_match) * 100 64.8% of unique dogs in the impound data, identified by their animal id number, have a different breed listed in the registration data. In many cases, this appears to reflect thedifficulty in identifying breeds visually, hence why many pit bull type breeds are grouped under ‘Pit Bull Terrier’.
But there are also many cases of dogs which appear visually distinct having mismatched breeds between the impound and registration data.
<- comparison_by_id |> filter (! is_match) |> count (breed_reg, breed_imp, sort = TRUE ) |> rename (reg_says = breed_reg,impound_says = breed_imp,frequency = nhead (top_mismatches, n = 10 )
# A tibble: 10 × 3
reg_says impound_says frequency
<chr> <chr> <int>
1 American Pit Bull Terrier Pit Bull Terrier 886
2 Fox (Smooth) Terrier Fox Terrier (Smooth) 64
3 Staffordshire Bull Terrier Pit Bull Terrier 61
4 American Staffordshire Terrier Pit Bull Terrier 36
5 American Staffordshire Terrier Staffordshire Bull Terrier 33
6 Staffordshire Bull Terrier American Staffordshire Terrier 32
7 Labrador Retriever Pit Bull Terrier 30
8 Labrador Retriever Staffordshire Bull Terrier 28
9 American Pit Bull Terrier Staffordshire Bull Terrier 27
10 Labrador Retriever Mastiff 25
Top 10 primary breeds as at FY2025:
<- registration |> filter (sheet_name == "FY25" ) |> calc_proportions ("breed_standardised" , count_name = "primary_breed" )|> rename ("Breed" = breed_standardised,"Count" = primary_breed,"Proportion" = prop|> mutate (Proportion = scales:: percent (Proportion, accuracy = 0.1 ),Count = format (Count, big.mark = "," )|> head (n = 10 ) |> kable () |> kable_styling () Dog population secondary breed counts
<- registration |> filter (sheet_name == "FY25" ) |> calc_proportions ("animal_breed2_description" , count_name = "secondary_breed" )head (secondary_breed, n = 10 )
# A tibble: 10 × 3
animal_breed2_description secondary_breed prop
<chr> <int> <dbl>
1 <NA> 75366 0.575
2 Cross 19253 0.147
3 Poodle, Miniature 3190 0.0243
4 Poodle, Standard 2885 0.0220
5 Retriever, Labrador 2861 0.0218
6 Poodle, Toy 1802 0.0137
7 Terrier, Staffordshire Bull 1789 0.0136
8 Spaniel, Cavalier King Charles 1519 0.0116
9 Shih Tzu 1501 0.0114
10 Collie, Border 1393 0.0106
Primary breeds most commonly classified as ‘menacing’ due to behaviour
<- |> filter (sheet_name == "FY25" ) |> filter (classification == "Menacing - observed or reported behaviour of dog" ) |> count (breed_standardised, name = "menacing" ) |> left_join (primary_breed |> rename (primary_prop = prop), by = join_by (breed_standardised)) |> filter (primary_breed > 100 ) |> mutate (menacing_prop = menacing / primary_breed) |> arrange (desc (menacing_prop)) head (menacing_breed, n = 10 )
# A tibble: 10 × 5
breed_standardised menacing primary_breed primary_prop menacing_prop
<chr> <int> <int> <dbl> <dbl>
1 Akita 8 102 0.000778 0.0784
2 Siberian Husky 63 1330 0.0101 0.0474
3 American Bulldog 66 1707 0.0130 0.0387
4 Neapolitan Mastiff 11 293 0.00223 0.0375
5 Mastiff 57 1526 0.0116 0.0374
6 Shar Pei 71 2445 0.0186 0.0290
7 Dogue de Bordeaux 4 139 0.00106 0.0288
8 Weimaraner 4 152 0.00116 0.0263
9 Bull Terrier 16 666 0.00508 0.0240
10 Bull Mastiff 24 1077 0.00821 0.0223
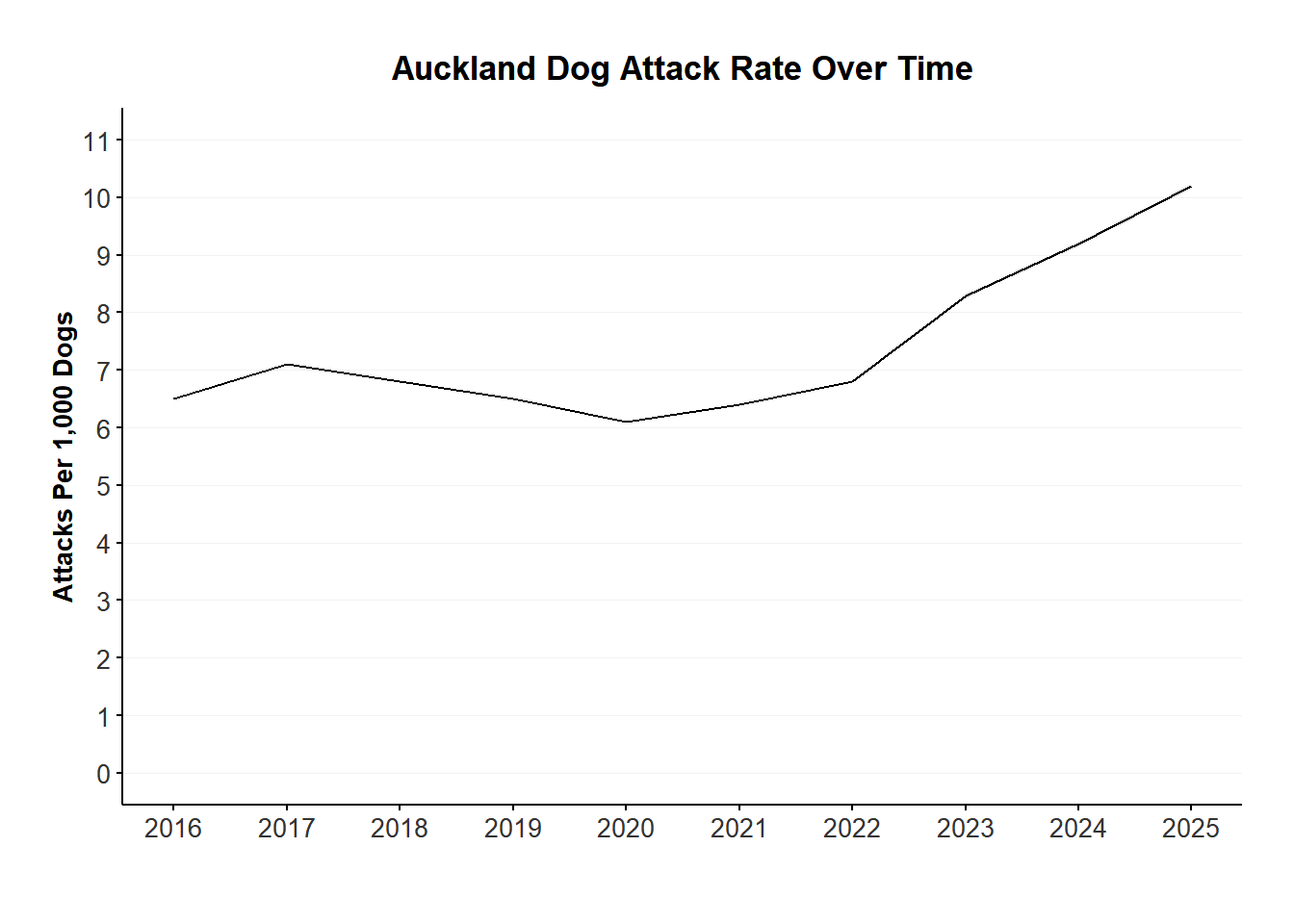
The rate of attacks on people per 1,000 dogs has increased from 6 in FY2016, to 10 in FY2025.
<- $ service |> filter (category == "attack-people" ) |> select (- category) |> rename (count_attacks = count) |> left_join ($ ownership |> filter (category == "dogs" ) |> select (- category) |> rename (count_dogs = count),by = join_by (year)) |> mutate (attack_rate = round ((count_attacks / count_dogs) * 1000 , digits = 1 ),head (attack_rate, n = 10 )
# A tibble: 10 × 4
year count_attacks count_dogs attack_rate
<dbl> <dbl> <dbl> <dbl>
1 2025 1341 131123 10.2
2 2024 1253 135546 9.2
3 2023 1098 131795 8.3
4 2022 848 125016 6.8
5 2021 756 118552 6.4
6 2020 685 112530 6.1
7 2019 716 110969 6.5
8 2018 745 110012 6.8
9 2017 816 115544 7.1
10 2016 740 114519 6.5
ggplot (attack_rate |> filter (year > 2015 ), aes (x = year, y = attack_rate)) + geom_line () + scale_y_continuous (limits = c (0 , 11 ), breaks = seq (0 , 11 , by = 1 )) + scale_x_continuous (breaks = 2016 : max (attack_rate$ year)) + xlab ("" ) + ylab ("Attacks Per 1,000 Dogs" ) + plot_theme () + labs (title = "Auckland Dog Attack Rate Over Time" ) Impounds
<- $ ownership |> filter (category == "dogs" ) |> rename ("dogs" = "count" ) |> select (- category) |> left_join (report_data$ impounds |> filter (category == "impounds" ),by = join_by (year)) |> rename ("impounds" = "count" ) |> select (- category) |> mutate (impound_rate = round ((impounds / dogs) * 1000 , digits = 1 )|> rename ("Year" = year,"Known Dogs" = dogs,"Impounded Dogs" = impounds,"Impound Rate" = impound_rate|> mutate (across (c (` Known Dogs ` , ` Impounded Dogs ` ), ~ format (.,big.mark = "," ))|> kable () |> kable_styling () Look at repeat impounds
<- impounds |> drop_na (dog_id) |> group_by (dog_id) |> add_count (name = "impounded_n" ) |> ungroup () |> select (dog_id, impounded_n, impound_date, impound_reason, status, suburb, desexed, clean_name) |> group_by (dog_id) |> reframe (impounded_n = impounded_n,reason = impound_reason,breed = clean_name|> arrange (desc (impounded_n)) |> select (dog_id, impounded_n, breed) |> unique ()|> count (impounded_n) |> mutate (proportion = n / sum (n))
# A tibble: 13 × 3
impounded_n n proportion
<int> <int> <dbl>
1 1 5452 0.755
2 2 1122 0.155
3 3 352 0.0487
4 4 155 0.0215
5 5 50 0.00692
6 6 33 0.00457
7 7 20 0.00277
8 8 24 0.00332
9 9 3 0.000415
10 10 5 0.000692
11 12 2 0.000277
12 18 2 0.000277
13 20 1 0.000138
Top 3 primary breeds for each attack severity rating for impounded dogs in FY2024
<- |> filter (impound_date > "2023-07-01" & ! is.na (severity)) |> group_by (severity) |> count (breed_standardised) |> arrange (desc (severity), desc (n))|> group_by (severity) |> slice_max (order_by = n, n = 3 ) |> arrange (desc (severity))
# A tibble: 19 × 3
# Groups: severity [6]
severity breed_standardised n
<chr> <chr> <int>
1 5 Pit Bull Terrier 17
2 5 Staffordshire Bull Terrier 12
3 5 American Bulldog 8
4 4 Pit Bull Terrier 30
5 4 Staffordshire Bull Terrier 25
6 4 American Bulldog 12
7 4 Shar Pei 12
8 3 Pit Bull Terrier 26
9 3 Staffordshire Bull Terrier 17
10 3 American Staffordshire Terrier 9
11 2 Pit Bull Terrier 25
12 2 Staffordshire Bull Terrier 17
13 2 Mastiff 10
14 1 Staffordshire Bull Terrier 10
15 1 American Bulldog 4
16 1 Pit Bull Terrier 4
17 0 Pit Bull Terrier 34
18 0 Staffordshire Bull Terrier 24
19 0 Labrador Retriever 9
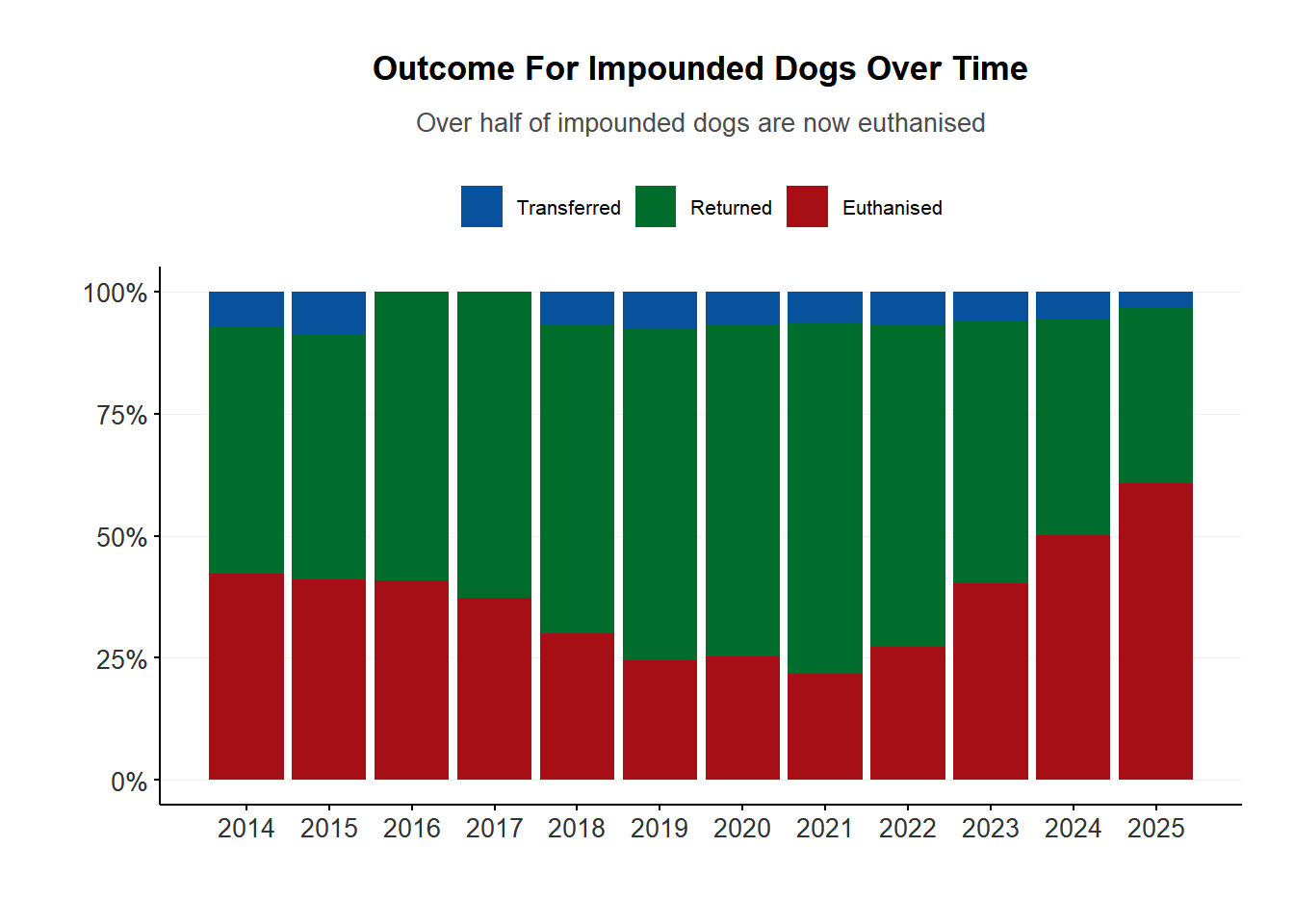
The overall rate of euthanising impounded dogs was dropping between 2014-2019. But after COVID, the rate has increased to be higher than ever before.
<- $ impounds |> filter (category != "impounds" )$ category <- factor (impound_result$ category, levels = c ("transferred" , "returned" , "euthanised" ))ggplot (impound_result, aes (x = year, y = count, fill = category)) + geom_bar (position = "fill" , stat = "identity" ) + scale_fill_manual (values = c ("#08519c" , "#006d2c" , "#a50f15" ),labels = tools:: toTitleCase) + scale_y_continuous (labels = scales:: label_percent ()) + scale_x_continuous (breaks = unique (impound_result$ year)) + xlab ("" ) + ylab ("" ) + plot_theme () + labs (title = "Outcome For Impounded Dogs Over Time" ,subtitle = "Over half of impounded dogs are now euthanised" )
Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_bar()`).
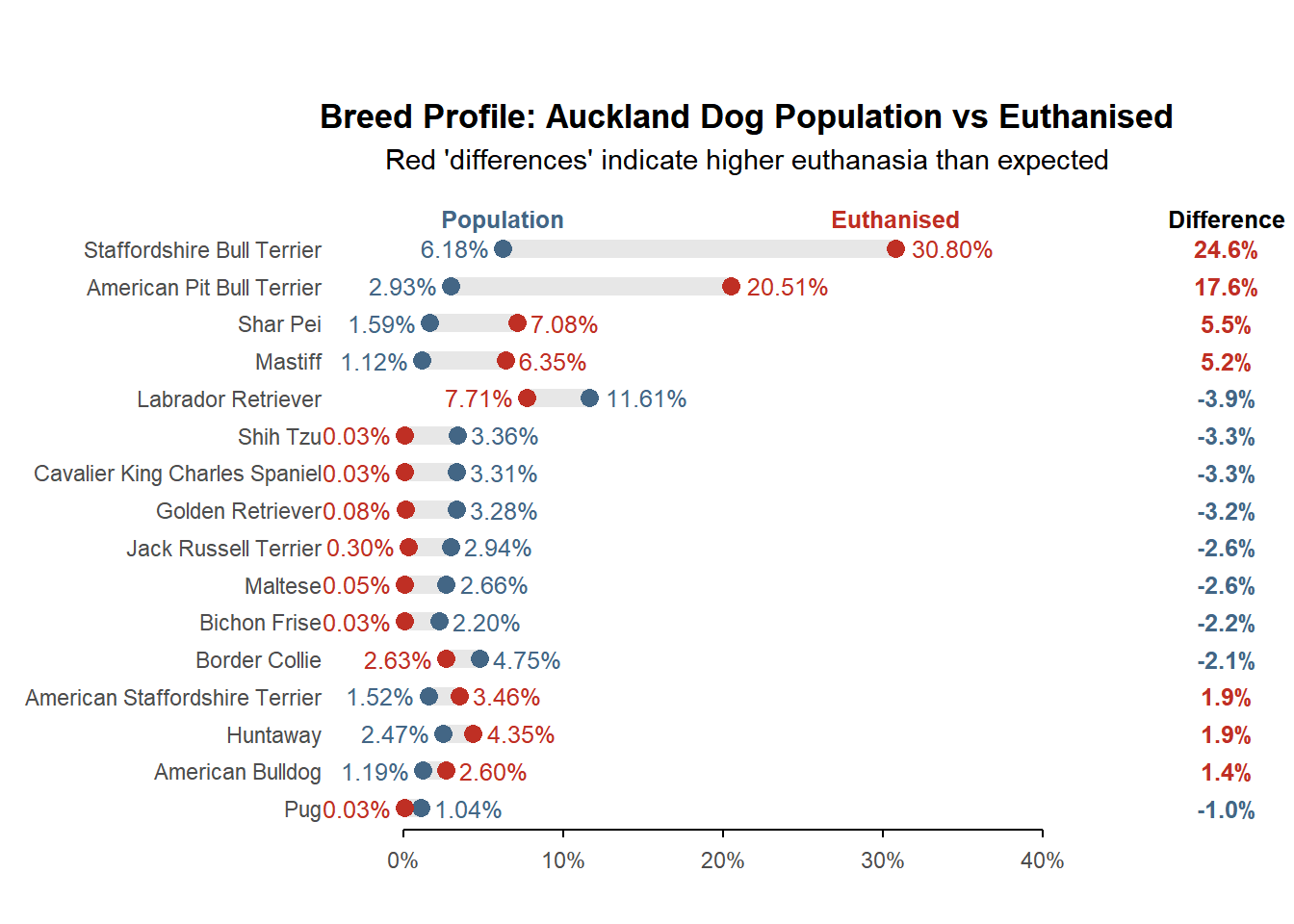
Impound outcome by primary breed (population and impound data are both FY2024)
<- registration |> filter (sheet_name == "FY24" ) |> count (primary_breed = breed_standardised, name = "n_pop" ) |> mutate (prop_pop = n_pop / sum (n_pop)) |> inner_join (|> filter (impound_date > "2023-07-01" , status %in% c ("Euthanised" , "For Euthanasia" )) |> mutate (primary_breed = str_replace (primary_breed, ".*Pit Bull Terrier.*" , "American Pit Bull Terrier" )) |> count (primary_breed, name = "n_euth" ) |> mutate (prop_euth = n_euth / sum (n_euth)),by = "primary_breed" |> mutate (diff = prop_euth - prop_pop,max_val = pmax (prop_euth, prop_pop) |> filter (abs (diff) > 0.01 ) |> arrange (abs (diff)) |> mutate (primary_breed = fct_inorder (primary_breed))<- c ("prop_euth" = "#BF2F24" , "prop_pop" = "#436685" )<- nrow (breed_data)<- breed_data |> pivot_longer (cols = starts_with ("prop" ), names_to = "type" , values_to = "val" ) |> ggplot (aes (val, primary_breed)) + geom_line (aes (group = primary_breed), color = "#E7E7E7" , linewidth = 3.5 ) + geom_point (aes (color = type), size = 3 ) + geom_text (aes (label = percent (val, 0.01 ), color = type, hjust = if_else (val == max_val, - 0.2 , 1.2 )), size = 3.25 ) + geom_text (data = \(.x) filter (.x, primary_breed == last (primary_breed)),aes (label = if_else (type == "prop_pop" , "Population" , "Euthanised" ), color = type),nudge_y = 0.8 , fontface = "bold" , size = 3.25 ) + scale_x_continuous (breaks = seq (0 , 0.4 , by = 0.10 ),labels = \(x) if_else (round (x, 2 ) %in% round (seq (0 , 0.4 , 0.1 ), 2 ), percent (x, accuracy = 1 ), "" ),guide = guide_axis (cap = "both" )+ scale_color_manual (values = colours) + coord_cartesian (xlim = c (- 0.02 , 0.45 ), ylim = c (1 , n_breeds + 1 ), clip = "off" ) + labs (title = "Breed Profile: Auckland Dog Population vs Euthanised" ,subtitle = "Red 'differences' indicate higher euthanasia than expected" , x = NULL , y = NULL ) + theme_minimal () + theme (legend.position = "none" , panel.grid = element_blank (), plot.title = element_text (face = "bold" , hjust = 0.5 ), plot.subtitle = element_text (hjust = 0.5 ),axis.line.x = element_line (color = "black" , linewidth = 0.5 ),axis.ticks.x = element_line (color = "black" , linewidth = 0.5 ),axis.ticks.length.x = unit (3 , "pt" )<- ggplot (breed_data, aes (x = 0 , y = primary_breed)) + geom_text (aes (label = percent (diff, 0.1 ), color = if_else (diff > 0 , "prop_euth" , "prop_pop" )),fontface = "bold" , size = 3.25 ) + annotate ("text" , x = 0 , y = n_breeds + 0.8 , label = "Difference" , fontface = "bold" , size = 3.25 ) + scale_color_manual (values = colours) + coord_cartesian (ylim = c (1 , n_breeds + 1 ), clip = "off" ) + theme_void () + theme (legend.position = "none" )+ p2 + plot_layout (widths = c (10 , 1 )) & theme (plot.margin = margin (20 , 5 , 10 , 5 )) Reasons for euthanasia
<- |> filter (impound_date > "2023-07-01" & ! is.na (euthanasia_reason)) |> mutate (euthanasia_reason = case_when (str_detect (euthanasia_reason, "Health|Parvo" ) ~ "Health or Parvo Virus" ,str_detect (euthanasia_reason, "Dog vs Dog|Dog vs Domestic|Dog vs Stock|Dog vs Poultry" ) ~ "S.57 Dog vs Animal" ,.default = euthanasia_reason|> group_by (euthanasia_reason) |> count (name = "count" ) |> ungroup () |> mutate (proportion = count / sum (count)|> arrange (desc (proportion))
# A tibble: 13 × 3
euthanasia_reason count proportion
<chr> <int> <dbl>
1 Failed Temperament Test 1963 0.501
2 Shelter Full 948 0.242
3 Health or Parvo Virus 518 0.132
4 S.57 Dog vs Animal 120 0.0306
5 S.57 Dog vs Person 106 0.0271
6 No Longer Suitable For Adoption 86 0.0219
7 S.57A Aggressive Behaviour 71 0.0181
8 Dependant Pup 37 0.00944
9 Relinquish - Aggressive Behaviour 31 0.00791
10 Destruction Order - S.64 19 0.00485
11 Menacing Dog - S.33A Deed 13 0.00332
12 Dangerous Dog - S.31 3 0.000766
13 Menacing Dog - S.33C Breed 3 0.000766
Total infringements skyrocket in 2025
$ enforcement |> filter (category == "infringements-total" )
# A tibble: 12 × 3
year category count
<dbl> <chr> <dbl>
1 2025 infringements-total 17430
2 2024 infringements-total 6387
3 2023 infringements-total 4748
4 2022 infringements-total 3271
5 2021 infringements-total 5126
6 2020 infringements-total 3480
7 2019 infringements-total 5172
8 2018 infringements-total 5817
9 2017 infringements-total 5098
10 2016 infringements-total 3835
11 2015 infringements-total 4521
12 2014 infringements-total 5638
The bulk of infringements are for failing to register a dog
$ enforcement |> filter (category == "failure-register" )
# A tibble: 12 × 3
year category count
<dbl> <chr> <dbl>
1 2025 failure-register 10149
2 2024 failure-register 2305
3 2023 failure-register 1490
4 2022 failure-register 763
5 2021 failure-register 1903
6 2020 failure-register 1138
7 2019 failure-register 2028
8 2018 failure-register 2534
9 2017 failure-register 2109
10 2016 failure-register 1496
11 2015 failure-register 2200
12 2014 failure-register 3203
But prosecutions remain at low levels
$ enforcement |> filter (category == "prosecutions-appeals" )
# A tibble: 12 × 3
year category count
<dbl> <chr> <dbl>
1 2025 prosecutions-appeals 141
2 2024 prosecutions-appeals 117
3 2023 prosecutions-appeals 121
4 2022 prosecutions-appeals 117
5 2021 prosecutions-appeals 134
6 2020 prosecutions-appeals 161
7 2019 prosecutions-appeals 220
8 2018 prosecutions-appeals 237
9 2017 prosecutions-appeals 217
10 2016 prosecutions-appeals 197
11 2015 prosecutions-appeals 137
12 2014 prosecutions-appeals 210
Auckland Dog Population By Region
Get Auckland human population by region
# Source: https://figure.nz/chart/bSr6yQmn9V9BFrXK <- read_csv ("../data/2023-auckland-population.csv" ) |> filter (` Census year ` == "2023" & ` Local board area ` != "Auckland" ) |> mutate (local_board = paste0 (` Local board area ` , " Local Board Area" ),local_board = case_when (== "Auckland Local Board Area" ~ "Total Auckland" ,.default = local_board)|> rename (census_year = ` Census year ` ,population = Value) |> select (census_year, local_board, population) |> left_join (locality |> select (local_board, region) |> unique (),by = join_by (local_board)) |> group_by (region) |> reframe (human_population = sum (population)
Rows: 154 Columns: 8
── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
chr (6): Census year, Local board area, Local board area Code, Measure, Valu...
dbl (1): Value
lgl (1): Null Reason
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.
Known dogs
<- |> filter (sheet_name == "FY25" ) |> left_join (locality, by = join_by (suburb)) |> select (suburb, local_board, region) |> group_by (suburb) |> add_count (suburb, name = "known_suburb" ) |> group_by (local_board) |> add_count (local_board, name = "known_board" ) |> group_by (region) |> add_count (region, name = "known_region" ) |> unique () |> arrange (desc (known_suburb)) |> ungroup ()|> select (region, known_region) |> distinct () |> left_join (auck_pop,by = join_by (region)) |> arrange (desc (known_region)) |> mutate (dogs_per_1k_human = (known_region / human_population) * 1000 ,across (c (known_region, human_population), label_comma ()),region = tools:: toTitleCase (region)|> rename ("Region" = region,"Dogs" = known_region,"People" = human_population,"Dogs Per 1,000 People" = dogs_per_1k_human|> kable () |> kable_styling () Dog population trends in each Local Board
<- |> select (sheet_name, suburb) |> left_join (locality, by = join_by (suburb)) |> group_by (sheet_name, local_board) |> count () |> pivot_wider (names_from = sheet_name, values_from = n) |> mutate (change = (FY25 - FY21) / FY21|> filter (FY25 > 100 ) |> arrange (desc (change)) |> ungroup ()|> mutate (across (contains ("FY" ), comma),change = scales:: percent (change, accuracy = 0.1 )|> rename ("2021" = "FY21" ,"2022" = "FY22" ,"2023" = "FY23" ,"2024" = "FY24" ,"2025" = "FY25" ,"Change" = "change" ,"Local Board" = "local_board" |> kable () |> kable_styling ()
median (dogs_board$ change, na.rm = TRUE ) While the median increase across Auckland local boards was 9%, the Interquartile Range (IQR) analysis below identifies Māngere-Ōtāhuhu and Ōtara-Papatoetoe Local Boards as statistical outliers for their significantly higher growth, distinct from the regional trend.
# Calculate upper fence <- dogs_board |> summarise (q1 = quantile (change, 0.25 ),q3 = quantile (change, 0.75 ),iqr = q3 - q1,upper_fence = q3 + (1.5 * iqr)# Flag high outliers <- dogs_board |> mutate (is_high_outlier = change > stats$ upper_fence,|> filter (is_high_outlier)
# A tibble: 2 × 8
local_board FY21 FY22 FY23 FY24 FY25 change is_high_outlier
<chr> <int> <int> <int> <int> <int> <dbl> <lgl>
1 Māngere-Ōtāhuhu Local Bo… 3407 3677 4173 4670 4809 0.412 TRUE
2 Ōtara-Papatoetoe Local B… 3912 4267 4668 5107 5192 0.327 TRUE
Registration data
<- |> filter (sheet_name == "FY25" ) |> mutate (current_reg = case_when (registration_latest_year < 2024 ~ FALSE , .default = TRUE ),owner_reg = case_when (owner_registration_class == "RDOL" ~ TRUE ),classification = case_when (classification == "Unknown" ~ FALSE ,str_detect (classification, "Menacing|Dangerous" ) ~ TRUE )|> left_join (regions_dogs, by = join_by (suburb)) |> select (current_reg, owner_reg, breed_standardised, animal_desexed, classification, suburb, local_board: known_region) <- |> pivot_longer (cols = c (current_reg, owner_reg, animal_desexed, classification),names_to = "metric" ,values_to = "is_true" |> group_by (suburb, local_board, region, metric) |> summarise (count = sum (is_true, na.rm = TRUE ),known_suburb = first (known_suburb),known_board = first (known_board),known_region = first (known_region),.groups = "drop" |> group_by (metric) |> mutate (p_suburb = count / known_suburb|> group_by (local_board, metric) |> mutate (p_board = sum (count) / first (known_board)) |> group_by (region, metric) |> mutate (p_region = sum (count) / first (known_region)) |> ungroup ()write_csv (regions_reg, "../outputs/regions_reg.csv" ) Compare Local Boards
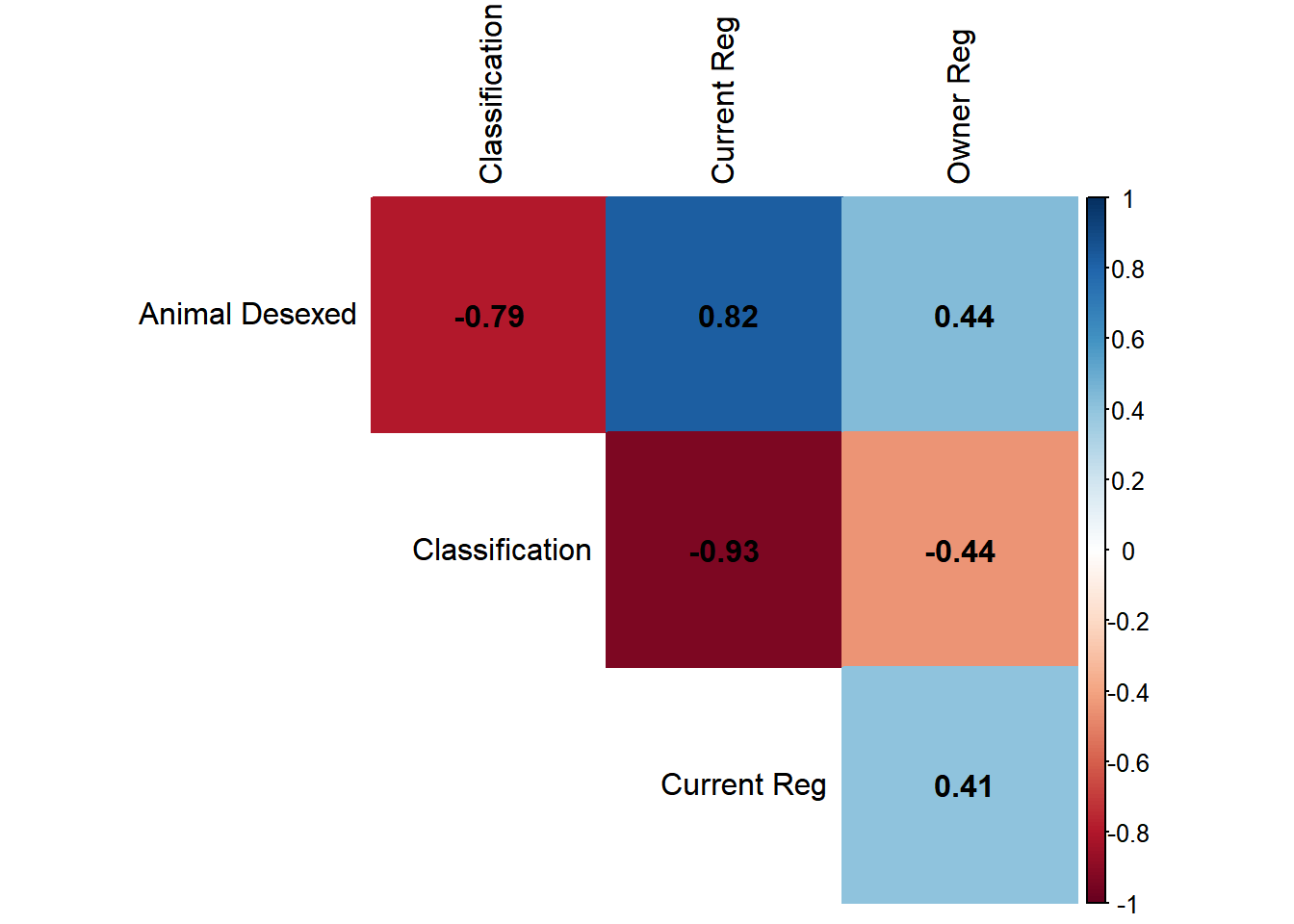
<- regions_reg |> select (local_board, metric, p_board) |> distinct () |> mutate (metric = metric |> str_replace_all ("_" , " " ) |> str_to_title (),local_board = local_board |> str_replace_all ("_" , " " ) |> str_to_title ()<- plot_data |> group_split (metric)<- plots_list |> reduce (\(p_obj, data_slice) {|> add_bars (data = data_slice,x = ~ p_board,y = ~ fct_reorder (local_board, p_board),name = unique (data_slice$ metric),marker = list (color = region_colours[which (plots_list == list (data_slice))]),visible = (identical (data_slice, plots_list[[1 ]])),orientation = 'h' ,showlegend = FALSE ,hovertemplate = "Board: %{y}<br>Value: %{x:.1%}<extra></extra>" .init = plot_ly (height = 800 ))<- plots_list |> imap (\(data_slice, idx) {<- rep (FALSE , length (plots_list))<- TRUE list (method = "restyle" ,args = list ("visible" , as.list (vis_vector)),label = unique (data_slice$ metric)<- p |> layout (title = list (text = "Animal Registration Metrics by Local Board (FY25)" ,x = 0.05 , y = 0.98 , xanchor = 'left' ,yanchor = 'top' ,font = list (size = 20 )margin = list (l = 180 , t = 100 , r = 30 , b = 50 ),xaxis = list (title = "" , tickformat = ".0%" ),yaxis = list (title = "" ),updatemenus = list (list (buttons = buttons,direction = "down" ,showactive = TRUE ,x = 0.10 , y = 1.05 , xanchor = 'left' ,yanchor = 'top' ,bgcolor = "#f8f9fa" :: saveWidget (interactive_plot, "../outputs/reg_metrics.html" , selfcontained = TRUE ) The below Spearman rank correlation analysis shows that Local Boards with high current registrations tend to have high rates of desexing and low rates of classified animals, while those with high rates of classifications tend to have lower rates of desexing.
<- plot_data |> pivot_wider (names_from = metric, values_from = p_board) |> select (- local_board) |> as.matrix ()<- rcorr (corr, type = "spearman" )corrplot (cor_results$ r, method = "color" , type = "upper" , addCoef.col = "black" , tl.col = "black" , diag = FALSE ,sig.level = 0.05 , insig = "blank" ) Attacks per 1,000 known dogs in each Local Board
<- rfs |> filter (>= "2024-07-01" ,str_to_lower (rfs_type) %in% c ("dog attack on people" , "dog attack on child" )|> count (suburb, name = "attacks_suburb" ) |> inner_join (regions_dogs, by = "suburb" ) |> group_by (local_board) |> mutate (attacks_board = sum (attacks_suburb)) |> group_by (region) |> mutate (attacks_region = sum (attacks_suburb)) |> ungroup () |> mutate (rate_suburb = (attacks_suburb / known_suburb) * 1000 ,rate_board = (attacks_board / known_board) * 1000 ,rate_region = (attacks_region / known_region) * 1000 |> select (suburb, local_board, region, starts_with ("attacks" ), starts_with ("known" ), starts_with ("rate" ))write_csv (regions_attacks, "../outputs/regions_attacks.csv" ) Primary breed profile for each region
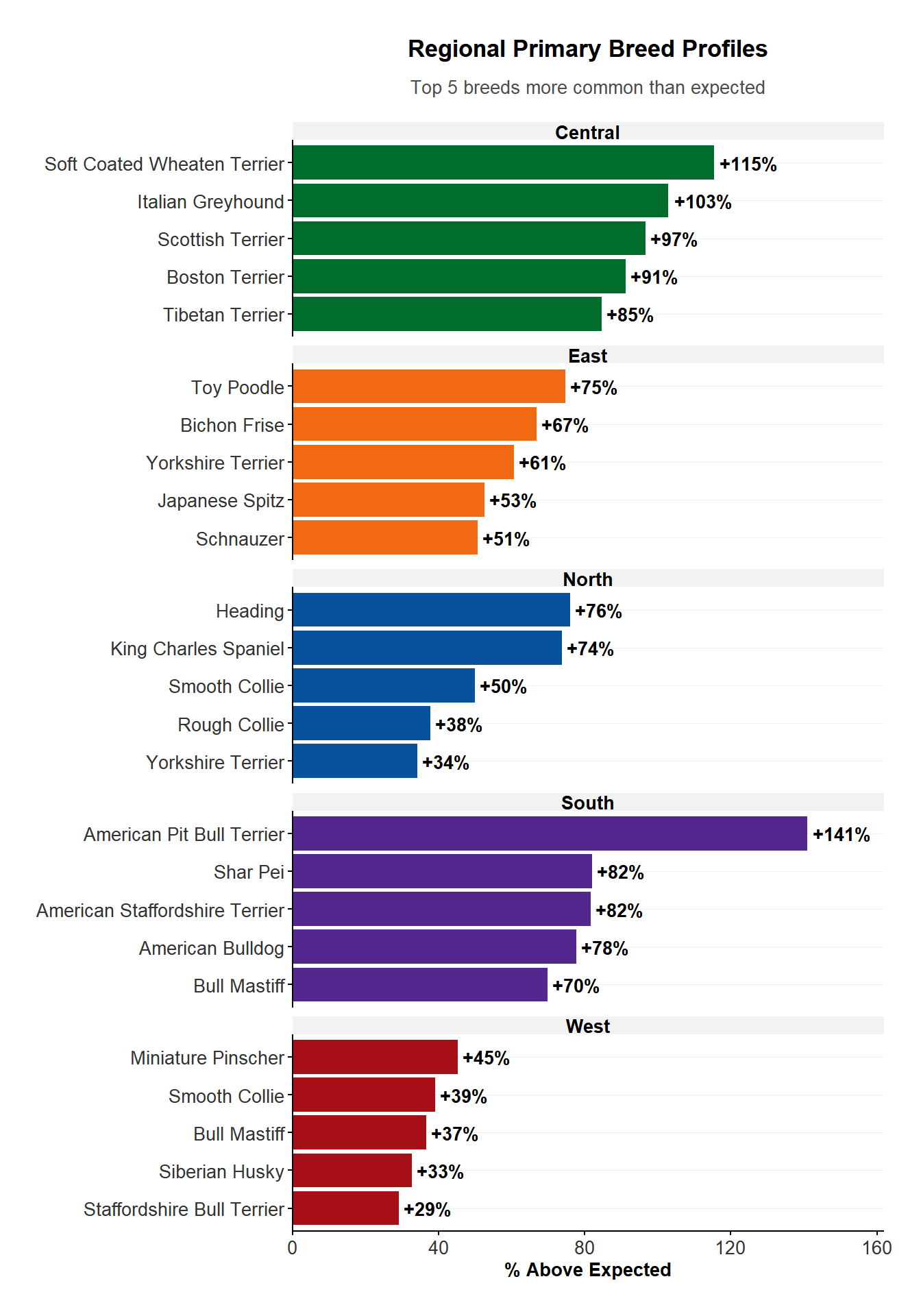
<- |> filter (sheet_name == "FY25" ) |> left_join (locality, by = join_by (suburb)) |> group_by (region) |> count (breed_standardised, name = "count" ) |> arrange (desc (count), .by_group = TRUE ) |> left_join (regions_dogs |> select (region, known_region) |> unique (), by = join_by (region)) |> mutate (prop_breed = format (count / known_region, scientific = FALSE ) Compare regional breed profiles statistically
<- regions_breed |> perform_chisq (group_var = "region" , cat_var = "breed_standardised" , count_var = "count"
Warning: Use of .data in tidyselect expressions was deprecated in tidyselect 1.2.0.
ℹ Please use `all_of(var)` (or `any_of(var)`) instead of `.data[[var]]`
write_csv (breeds_region, "../outputs/breed_over.csv" )
|> group_by (group) |> slice_max (percent_diff, n = 5 ) |> ungroup () |> ggplot (aes (x = reorder_within (category, percent_diff, group), y = percent_diff, fill = group)) + geom_col () + scale_fill_manual (values = region_colours) + geom_text (aes (label = paste0 ("+" , round (percent_diff), "%" )), hjust = - 0.1 , size = 3.5 , fontface = "bold" ) + scale_x_reordered () + coord_flip () + facet_wrap (~ group, scales = "free_y" , ncol = 1 ,labeller = function (labels) {lapply (labels, tools:: toTitleCase)+ scale_y_continuous (expand = expansion (mult = c (0 , 0.15 ))) + plot_theme () + theme (legend.position = "none" ) + labs (title = "Regional Primary Breed Profiles" ,subtitle = "Top 5 breeds more common than expected" ,x = NULL , y = "% Above Expected" Impounds per 1,000 dogs (FY2024 data)
<- impounds |> filter (impound_date > "2023-07-01" ) |> count (suburb, name = "impounds_suburb" ) |> inner_join (locality, by = "suburb" ) |> left_join (regions_dogs |> select (local_board, known_board, known_region), by = "local_board" ) |> unique () |> group_by (local_board) |> mutate (impounds_board = sum (impounds_suburb)) |> group_by (region) |> mutate (impounds_region = sum (impounds_suburb)) |> ungroup () |> mutate (rate_board = (impounds_board / known_board) * 1000 ,rate_region = (impounds_region / known_region) * 1000 |> select (suburb, local_board, region, starts_with ("impounds" ), starts_with ("known" ), starts_with ("rate" ))
Warning in left_join(inner_join(count(filter(impounds, impound_date > "2023-07-01"), : Detected an unexpected many-to-many relationship between `x` and `y`.
ℹ Row 1 of `x` matches multiple rows in `y`.
ℹ Row 23 of `y` matches multiple rows in `x`.
ℹ If a many-to-many relationship is expected, set `relationship =
"many-to-many"` to silence this warning.
write_csv (regions_impounds, "../outputs/regions_impounds.csv" ) Impound rate for attacks/rushing (FY2024 data)
<- c ("Relq - Sec 57" , "Sec 57" )<- c ("Sec 57A" )<- impounds |> filter (> "2023-07-01" ,%in% c (attack_codes, rush_codes)|> count (suburb, name = "impounds_attack_suburb" ) |> inner_join (locality, by = "suburb" ) |> left_join (regions_dogs |> select (local_board, known_board, known_region), by = "local_board" ) |> unique () |> group_by (local_board) |> mutate (impounds_attack_board = sum (impounds_attack_suburb, na.rm = TRUE )) |> group_by (region) |> mutate (impounds_attack_region = sum (impounds_attack_suburb, na.rm = TRUE )) |> ungroup () |> mutate (across (starts_with ("known" ), ~ replace_na (.x, 0 )),rate_attack_board = (impounds_attack_board / known_board) * 1000 ,rate_attack_region = (impounds_attack_region / known_region) * 1000 |> mutate (across (starts_with ("rate" ), ~ if_else (is.infinite (.x), NA_real_ , .x))) |> select (suburb, local_board, region, starts_with ("impounds" ), starts_with ("known" ), starts_with ("rate_attack" ))
Warning in left_join(inner_join(count(filter(impounds, impound_date > "2023-07-01", : Detected an unexpected many-to-many relationship between `x` and `y`.
ℹ Row 1 of `x` matches multiple rows in `y`.
ℹ Row 29 of `y` matches multiple rows in `x`.
ℹ If a many-to-many relationship is expected, set `relationship =
"many-to-many"` to silence this warning.
write_csv (regions_impounds_attacks, "../outputs/regions_impounds_attacks.csv" ) Outcome of impound by area
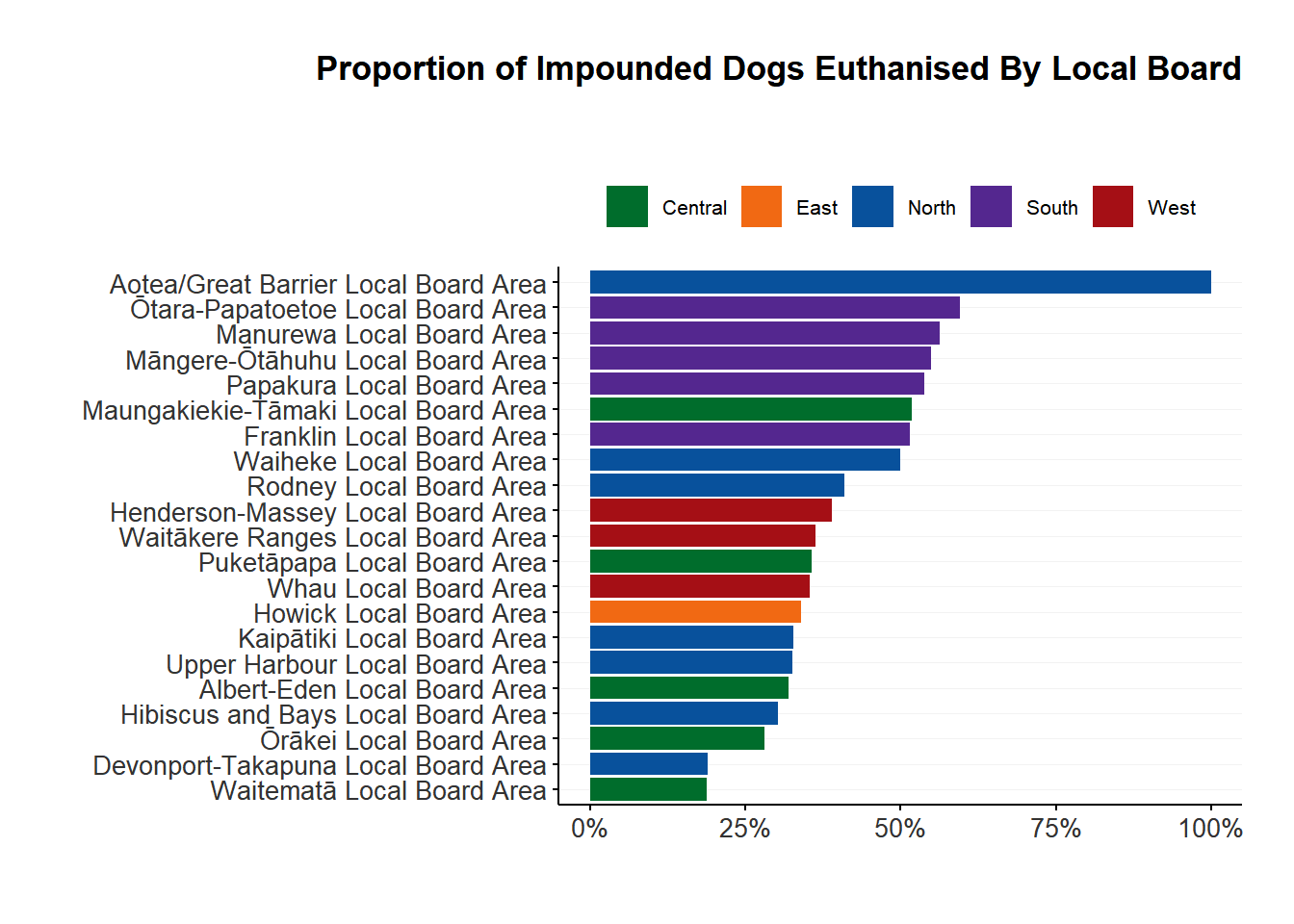
<- impounds |> filter (impound_date > "2023-07-01" ) |> mutate (status = case_match (status, "For Euthanasia" ~ "Euthanised" ,c ("Adoption Pending" , "Fostered" , "Hold - Adoption" ) ~ "Adopted" ,c ("Trfd to Breed Rescue" , "Trfd to Other TA" , "Trfd to Rescue Group" , "Trfd to SPCA" ) ~ "Rescued" ,c ("Dead on Arrival" , "Died in Shelter" , "Escaped" , "Hold - General" , "Hold - Legal Reason" , "Hold - Temp Test" , "Processing" , "Stolen" ) ~ "Other" ,.default = status|> left_join (regions_dogs, by = "suburb" ) |> count (region, local_board, status, name = "count" ) |> group_by (local_board) |> mutate (board_total = sum (count),proportion = count / board_total|> ungroup ()
Warning: There was 1 warning in `mutate()`.
ℹ In argument: `status = case_match(...)`.
Caused by warning:
! `case_match()` was deprecated in dplyr 1.2.0.
ℹ Please use `recode_values()` instead.
write_csv (regions_impounds_outcome, "../outputs/regions_impounds_outcome.csv" )|> filter (status == "Euthanised" & ! is.na (local_board)) |> ggplot (aes (x = fct_reorder (local_board, proportion), y = proportion, fill = region)) + geom_col () + coord_flip () + scale_fill_manual (values = region_colours,labels = stringr:: str_to_title) + scale_y_continuous (labels = scales:: label_percent ()) + plot_theme () + theme (plot.title = element_text (hjust = 1 )) + labs (title = "Proportion of Impounded Dogs Euthanised By Local Board" ,subtitle = "" ) + xlab ("" ) + ylab ("" ) Impounds by breed
<- impounds |> filter (impound_date > "2023-07-01" ) |> count (primary_breed, name = "impound_count" ) |> inner_join (|> select (- prop),by = join_by (primary_breed == breed_standardised)|> mutate (impound_rate = round ((impound_count / primary_breed.y) * 1000 , digits = 1 )) |> filter (primary_breed.y > 99 ) |> arrange (desc (impound_rate)) |> head (n = 10 )|> rename ("Breed" = primary_breed,"Impounds" = impound_count,"Population" = primary_breed.y,"Impound Rate" = impound_rate) |> mutate (across (c (` Impounds ` , ` Population ` ), ~ format (.,big.mark = "," ))|> kable () |> kable_styling ()